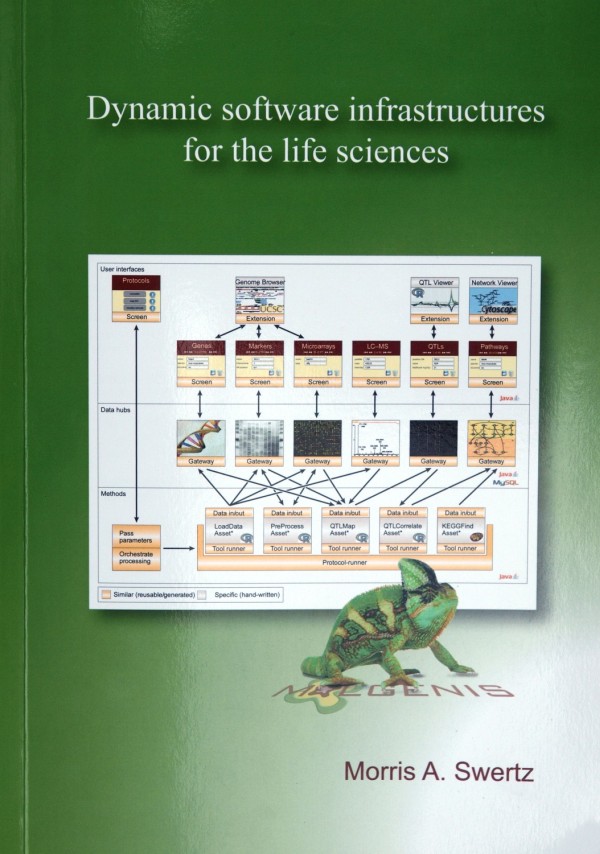
Progress in life sciences research has produced enormous amounts of data but the production of suitable software infrastructure to process these data has not been able to keep up. Part of the problem is that research groups each have their own needs for biotechnologies, protocols and specific species on which research is done.
Usually, bioinformaticians hand write the software code to adapt the existing software infrastructures to their needs. "Software developers have a luxury problem," says Swertz. "Virtually anything you want can be changed and adapted. However, doing this is extremely time consuming." Inevitably, the wheel is reinvented over and over again, he says. "It is fairly easy to copy reusable blocks of code from earlier systems into your own new program. But the system grows very large very quickly by doing this. Larger systems are exponentially more difficult and time consuming to maintain or update."
Blueprint
Therefore, Swertz has devised a simple programming language for these reusable blocks of code, a so-called Domain Specific Language (DSL). The domain in his case is biology. The 'specific' part means standardizing a number of operations. This may be an input box on the user interface or a database query to retrieve data. A bioinformatician setting up a system for example for a microarray experiment only has to tell the system, using DSL, that he needs this type of experiment. "It is the blueprint of your system," says Swertz.
Next, this short piece of code needs to be translated into a computer language that the system in use will understand. Swertz devised a computer programme called a generator which can do this. His generator, translates the short instructions of the bioinformatician into Java, the language his systems use. "The large advantage is that the generated Java code does not require maintenance. Should you wish to change the system you only need to change the blueprint and run the generator again. Additionally, if you wish to change the output Java code, you can do so. But keep in mind that it is extremely important that this handwritten code and the generated could are kept in separate folders."
The full toolbox of reusable blocks of code, the DSL programming language and the generator to convert DSL into a usable programming language is called Molgenis. It is unique in that it is not only a generator but a full toolbox. A number of other generators exist, each with its own specialty. Molgenis is specifically aimed at data management.
Synergy
Swertz already made a prototype of Molgenis before he even started as a PhD student in Groningen. He ran a company that sells tools for creating websites. Molgenis started as such a tool. The department of Molecular Genetics of Groningen University asked him to do a project for them and subsequently decided to offer him a PhD position. "The bioinformatics world is very dynamic, completely different from the insurance world I was used to. A kind of Wild West. I found it a lot more challenging and decided to stay." After a year of research the geneticists asked him for help on a specific project related to the one he did earlier. "They were still designing their experiments and we were already designing their data management system. This got to the point where we could not only make the software system but also help them organising their experiments. It was a great synergy."
The system itself was relatively simple. The lab needed to perform microarray experiments. The first step is designing the microarray plates. The second step is preparing their bacteria and applying these to the microarray. Finally, a picture is taken of the plates and analysed. Swertz' system automated the administration of all the data.
Since the first project in 2002, the Molgenis programme is technically completely changed. "The great thing is that it is very easy for life science researchers to update their systems with new technologies. Just run the updated generator with the blueprint," explains Swertz. New adaptations include the ability to handle much larger datasets. "We used to run data as a black box. Now we want to be able to analyze each step separately. This means the use of many different file types and many more separate sample data."
He even works on a Molgenis version which can not only generate new databases from an experiment but also import existing databases and compare data in a standardized way. The request comes from the European Casimir consortium that does genetic research on mice. "Casimir wanted to open up existing mice databases for future access. Bioinformaticians will be able to answer questions from researchers much faster."
Beyond borders
Swertz is looking beyond the borders of The Netherlands in more than one way. The Molecular Genetics group in Groningen where Swertz now works as a post-doc has a cooperation with a group in Manchester. This group devised the Taverna system which standardizes a workflow of operations by clicking on blocks of existing processors and linking input and output together. Integrating Molgenis and Taverna would give a user two functionalities in one, explains Swertz. "In practice you would for example see a Taverna world with blocks of operations." One of these blocks would be Molgenis. Conversely, the user interface of Molgenis may have a button saying 'calculate' which starts up Taverna. "We are both happy with this cooperation.They are good at something we are not, and we are good at something they are not."
Additionally, Swertz works on EU-wide standardization of the blueprint. "Standardization of the working system, the output of the generator is one thing. But research groups still need to define their own data structure, their blueprint. If the blueprint is inadequate, the generator output is useless. If we could set up a number of standard blueprints, for example a data model for genetical genomics (see box) as we are doing here in Groningen, research groups would not only be able to communicate on a technical level but also actually understand each other."
Dit artikel is gepubliceerd in het Engelstalige blad Interface nr. 1, 2008.